Introduction
AxonDeepSeg is an open-source software using deep learning and aiming at automatically segmenting axons and myelin sheaths from microscopy images. It performs multi-class semantic segmentation using a convolutional neural network.
AxonDeepSeg was developed at NeuroPoly Lab, Polytechnique Montreal, University of Montreal, Canada.
Quick Start
These instructions are for Linux/macOS. For Windows instructions, see the installation section. For a video demonstration, see below.
Download and install AxonDeepSeg (requires git):
git clone https://github.com/neuropoly/axondeepseg.git cd axondeepseg ./install_ads -y
Close and reopen your terminal to load the environment variables.
Launch the GUI:
ads_napariOpen the plugin: In Napari, go to
Plugins -> ADS (AxonDeepSeg)Segment an image: - Load your microscopy image via
File -> Open file(s)- Select the generalist model from the dropdown - Click Segment imageExtract morphometrics: Click the Morphometrics button to generate measurements for each axon.
For more detailed instructions, see the installation and usage sections below.
Help
Whether you are a newcomer or an experienced user, we will do our best to help and reply to you as soon as possible. Of course, please be considerate and respectful of all people participating in our community interactions.
If you encounter difficulties during installation and/or while using AxonDeepSeg, or have general questions about the project, you can start a new discussion on AxonDeepSeg GitHub Discussions forum. We also encourage you, once you’ve familiarized yourself with the software, to continue participating in the forum by helping answer future questions from fellow users!
If you encounter bugs during installation and/or use of AxonDeepSeg, you can open a new issue ticket on the AxonDeepSeg GitHub issues tracker.
Training Models
Training models requires a significant amount of quality data, high-performance computational resources, and most importantly–expertise. We don’t recommend you train, retrain, or finetune our models without first contacting the AxonDeepSeg team.
If the current models available in AxonDeepSeg do not perform sufficiently well with your dataset or for your applications and you are interested in training a model for AxonDeepSeg for your specific dataset, please reach out to the AxonDeepSeg team at axondeepseg@googlegroups.com to schedule a video-call to discuss more.
Note
To get more information on how our current models were trained, please refer to this GitHub repository: https://github.com/axondeepseg/model_seg_generalist
Testimonials
Do you also use AxonDeepSeg and would like to share your feedback with the community? Please add your testimonial by clicking here. Thank you 😊
Anne Wheeler, PhD | Hospital for Sick Children | Canada 🇨🇦
Our lab uses ADS to segment and extract morphometrics of myelinated axons from from EM images of mouse white matter tracts. We have two in-progress studies where ADS is allowing us to efficiently extract this data in the context of abberant white matter development. ADS is very well documented and easy to use and implement. In addition, the developers have been very responsive to our requests for additional functionality. Thank you!
Alison Wong, MD/MSE | Dalhousie University | Canada 🇨🇦
I found AxonDeepSeg very helpful for my research on peripheral nerve injury and regeneration. It performed well at segmentation and very well at obtaining the measurements, this greatly increased the ability to analyze out outcomes. There will always be attempts at new and better software, but the fact that the AxonDeepSeg team has focused on an open source format with continued development is commendable. I found the GitHub to be essential.
Osvaldo Delbono, PhD | Wake Forest University School of Medicine | United States 🇺🇸
We utilize AxonDeepSeg for post-mortem nerve analysis of individuals afflicted with Alzheimer’s Disease, related dementias, parkinsonism, and vascular deterioration affecting both the central and peripheral nervous systems. Given that our samples comprise thousands of axon/myelin units within the tibialis nerve, AxonDeepSeg is indispensable for our research. The documentation for AxonDeepSeg is comprehensive, with the guidelines for software installation being especially helpful.
Alan Peterson, PhD | McGill University | Canada 🇨🇦
Our investigation involved 6 lines of gene-edited mice that elaborate myelin sheaths of greatly different thickness. We used AxonDeepSeg to quantify myelin/axon relationships in tiled EM images from multiple tracts in young to old mice thus making this a very large experiment. AxonDeepSeg worked perfectly throughout. To obtain the maximum resolution we excluded fibers in which demonstrated fixation artifacts such as myelin splitting and the filtering step was easily accommodated in the work flow. Along the way, we required minimal support but when needed, it was both excellent an timely.
Installation
The following sections will help you install all the tools you need to run AxonDeepSeg.
Which installation method should I use?
What operating system are you using?
│
├── Linux/macOS
│ └── Install via GitHub (recommended)
│ • Automatic environment setup
│ • Runs tests to verify installation
│ • Easy updates via git pull
│
├── Windows
│ └── Install via GitHub (recommended)
│ • Automatic environment setup
│ • Runs tests to verify installation
│ • Requires additional environment variable configuration (see instructions)
│
└── Any OS with existing Python 3.11/3.12 environment
│
├── Do you already have Napari installed?
│ │
│ ├── YES → Install via Napari Plugin Manager
│ │ • Integrates with existing Napari setup
│ │
│ └── NO → Install via pip
│ • Installs AxonDeepSeg and Napari together
│
└── Both options:
• Require manual Python environment setup
• No automatic test verification
Summary:
GitHub (recommended): Best for most users on any OS. Handles environment setup automatically and verifies installation with tests.
pip: For users who already have a Python 3.11/3.12 environment configured.
Napari Plugin Manager: For users who already have Napari installed.
Install Options
To install AxonDeepSeg, in a terminal window (macOS or Linux) or Command Prompt (Windows), “clone” AxonDeepSeg’s repository (you will need to have git installed on your system) and then open the directory:
git clone https://github.com/neuropoly/axondeepseg.git
cd axondeepseg
Note
The installation script automatically runs tests at the end to verify proper installation.
For Linux or macOS systems, run:
./install_ads
and follow the prompts. Or, if you want to install AxonDeepSeg with default settings, run:
./install_ads -y
After the installation is complete, close and reopen your terminal to load the environment variables before using AxonDeepSeg commands.
For Windows systems, run:
install_ads.bat
Then, to use ADS’s command-line scripts in Command Prompt, please follow these instructions:
Open the Start Menu -> Type ‘edit environment’ -> Open ‘Edit environment variables for your account’
Click ‘New’, then enter ‘ADS_DIR’ for the variable name. For the value, copy and paste the path to the ADS directory (see full path displayed after install_ads.bat is completed)
Click ‘OK’, then click on the ‘Path’ variable, then click the ‘Edit…’ button.
Click ‘New’, then copy and paste the (ADS_PATH)bin directory:
Click ‘OK’ three times. You can now access ADS’s scripts in the Command Prompt.
Updating AxonDeepSeg
To update the AxonDeepSeg to the latest version, run:
git checkout master && git pull
and then follow the installation instructions above specified for your operating system.
Note
This installation method does not support running the test suite. For software development, please use the installation via git.
This method is only compatible with Python 3.11.x and 3.12.x.
Virtual environments are recommended. See the Python venv documentation or conda documentation for instructions.
This section provides instructions for installing the AxonDeepSeg plugin directly via pip. This method is useful if you prefer to manage your Python packages using pip instead of the Napari Plugin Manager.
Install AxonDeepSeg: Install the AxonDeepSeg plugin via pip by running the following command:
pip install "axondeepseg>=5"
This will install the latest version of AxonDeepSeg (version 5 or higher) along with its dependencies.
Launch Napari: After installation, launch Napari by running:
napari
Open the AxonDeepSeg Plugin: Once Napari is open, navigate to the top menu bar and click on:
Plugins -> ADS (AxonDeepSeg)
This will open the AxonDeepSeg plugin interface.
You’re ready to segment images!
Troubleshooting
For additional support, refer to the AxonDeepSeg documentation or the Napari Plugin Manager documentation.
AxonDeepSeg can also be installed via the Napari Plugin Manager. Install Napari following the Napari installation instructions. Then, use the Napari Plugin Manager to search for and install axondeepseg.
Note
AxonDeepSeg is only compatible with Python 3.11.x and 3.12.x. Ensure your Napari installation uses a compatible Python version.
For any issues related to Napari installation, please reach out to the Napari community via their GitHub issues.
Graphical User Interface (GUI)
AxonDeepSeg can be run via a Graphical User Interface (GUI) instead of the Terminal command line. This GUI is a plugin for the software Napari. Beyond the convenience of running AxonDeepSeg with the click of a button, this GUI is also an excellent way to manually correct output segmentations (if needed).
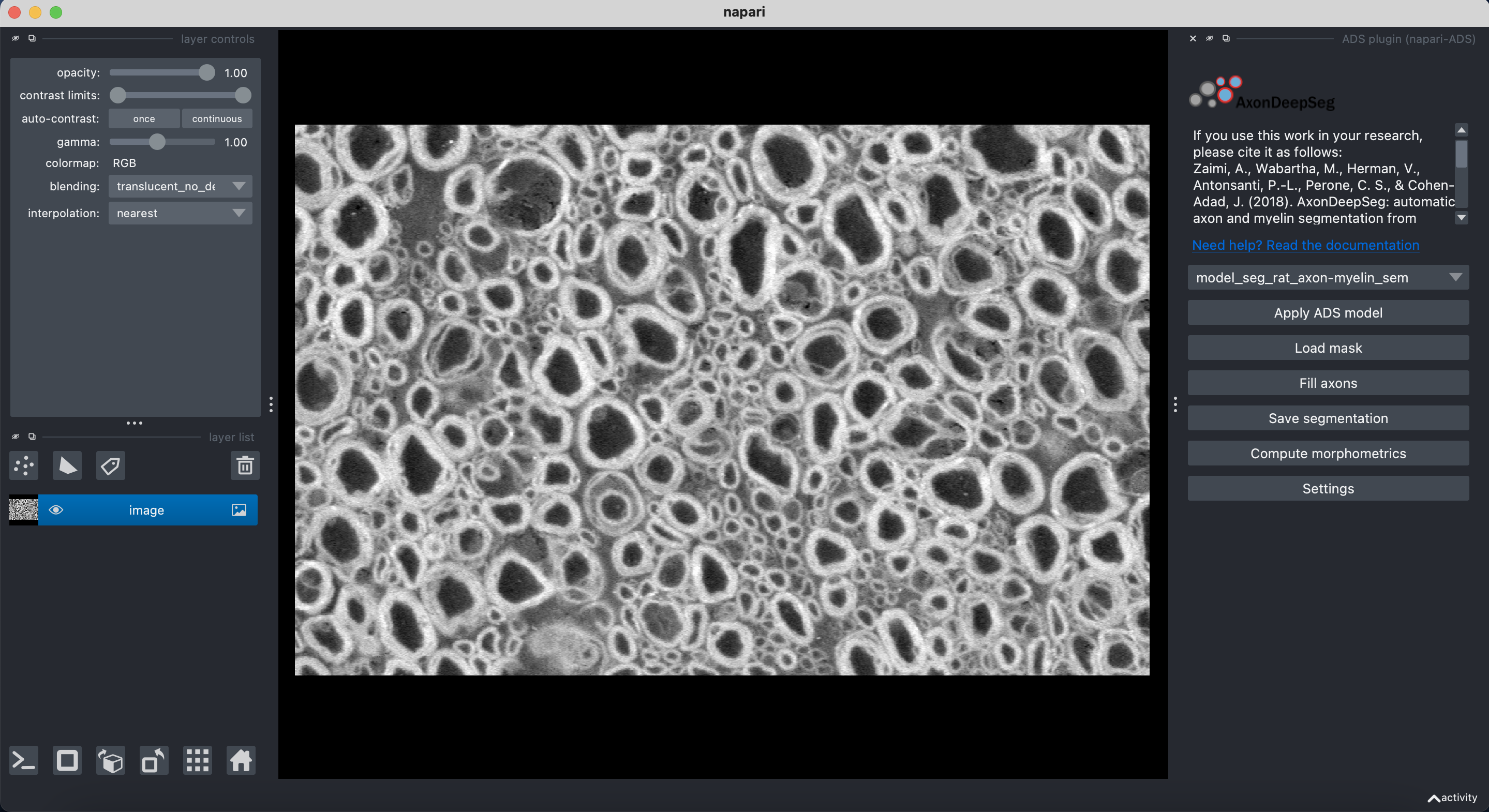
Launch Napari
ads_napari
In Napari, do the following:
- Click on Plugins -> ADS (AxonDeepSeg)
In the Manual correction of segmentation masks section, we provide a short tutorial describing how to use the AxonDeepSeg plugin for Napari.
Software Development with AxonDeepSeg
Software development with AxonDeepSeg requires an installation via git.
To develop code in the AxonDeepSeg software package and use it in a python shell, and to run the full test suite, you’ll need to first activate your virtual environment:
ads_activate
Note
To switch back to your default environment, run:
conda deactivate
If using AxonDeepSeg in a Jupyter Notebook, you’ll need to ensure that the virtual environment’s kernel is selected, or open Jupyter notebook from within an activated virtual environment.
Existing models
The generalist model is shipped together with the installation package and is recommended for initial use of the software.
Several other more specialized models are available.
For each model, a light and/or ensemble version is available. Light models are smaller size, whereas the ensemble versions may have slightly better segmentation performance.
Here are the details of all the models currently supported by AxonDeepSeg:
The following models are available for download:
Model |
Description |
|---|---|
generalist |
Multi-domain axon and myelin segmentation model trained on TEM, SEM, BF and CARS data. |
dedicated-BF |
Axon and myelin segmentation model trained on Bright-Field data. |
dedicated-SEM |
Axon and myelin segmentation model trained on Scanning Electron Microscopy data. |
dedicated-CARS |
Axon and myelin segmentation model trained on Coherent Anti-Stokes Raman Scattering data. |
unmyelinated-TEM |
Segments both myelinated and unmyelinated axons on TEM data. Also segments oligodendrocytes (nuclei and cytoplasmic processes). |
To download these models, you must first have AxonDeepSeg installed. Afterwards, use the following command to list available models:
download_model --list
Then, use the following to download a specific model:
download_model -m <model name>
Using AxonDeepSeg
Example dataset
You can demo the AxonDeepSeg by downloading the test data available here. It contains two SEM test samples and one TEM test sample.
Segmentation
Syntax
The script to launch is called axondeepseg. It takes several arguments:
Required arguments:
- -i IMGPATH
Path to the image to segment or path to the folder where the image(s) to segment is/are located.
Optional arguments:
- -m MODEL
Folder where the model is located, if different from the default model.
- -v VERBOSITY
Verbosity level.
0 (default): Quiet mode. Shows minimal information on the terminal.
1: Developer mode. Shows more information on the terminal, useful for debugging..
- --gpu-id GPU_ID
Number representing the GPU ID for segmentation if available. Default: None.
- --allow-large-images
Allow processing images that exceed PIL’s default decompression bomb pixel limit (~89 million pixels). Use this flag when segmenting very large microscopy images that would otherwise be rejected.
Note
You can get the detailed description of all the arguments of the axondeepseg command at any time by using the -h argument:
axondeepseg -h
Segment a single image
To segment a single microscopy image, specify the path to the image to segment in the -i argument. For instance, to segment the demo image used in our software tests (‘test/__test_files__/__test_demo_files__/image.png’), use the following command:
axondeepseg -i test/__test_files__/__test_demo_files__/image.png
The segmented acquisition will be saved in the same folder as the acquisition image, with the suffix ‘_seg-axonmyelin.png’, in png format, along with the binary axon and myelin segmentation masks (with the suffixes ‘_seg-axon.png’ and ‘_seg-myelin.png’). In our example, the following output files will be generated: image_seg-axonmyelin.png’, ‘image_seg-axon.png’ and ‘image_seg-myelin.png’.
Segment multiple images of the same resolution
To segment multiple microscopy images of the same resolution that are located in the same folder, specify the path to the folder in the -i argument. For instance, to segment the images in folder ‘test_sem_image/image1_sem/’, use the following command:
axondeepseg -i test_segmentation/test_sem_image/image1_sem/
Please note that when using axondeepseg, a file called axondeepseg.log will be saved in the current working directory. The console output will be saved in this file so you can review it later (useful to process large folders).
Segment images from multiple folders
To segment images that are located in different folders, specify the path to the folders in the -i argument, one after the other. For instance, to segment all the images of folders ‘test_sem_image/image1_sem/’ and ‘test_sem_image/image2_sem/’ of the test dataset, use the following command:
axondeepseg -i test_segmentation/test_sem_image/image1_sem/ test_segmentation/test_sem_image/image2_sem/
Morphometrics
You can generate morphometrics using AxonDeepSeg via the command line interface.
Syntax
The script to launch is called axondeepseg_morphometrics. It has several arguments.
Required arguments:
- -i IMGPATH
Path to the image file whose morphometrics needs to be calculated.
Optional arguments:
- -s SIZEPIXEL
Pixel size of the image(s) to segment, in micrometers. If no pixel size is specified, a pixel_size_in_micrometer.txt file needs to be added to the image folder path (that file should contain a single float number corresponding to the resolution of the image, i.e. the pixel size). The pixel size in that file will be used for the morphometrics computation.
- -a AXONSHAPE
Axon shape circle: Axon shape is considered as circle. In this case, diameter is computed using equivalent diameter. ellipse: Axon shape is considered as an ellipse. In this case, diameter is computed using ellipse minor axis. The default axon shape is set to circle.
- -f FILENAME
Name of the excel file in which the morphometrics will be stored. The excel file extension can either be .xlsx or .csv. If name of the excel file is not provided, the morphometrics will be saved as axon_morphometrics.xlsx.
- -c
Flag to save the colorized instance segmentation. For more information about this feature, see the Colorization subsection below.
- -u
Toggles unmyelinated mode. This will compute morphometrics for unmyelinated axons. Note that this requires a separate unmyelinated axon segmentation mask with suffix
_seg-uaxon.- -n
Computes morphometrics specific to nerve sections using the
-noption. This enables analysis of axons within nerve fascicle boundaries, based on a segmentation mask with the suffix_seg-nerve.png.- -d
Generate a diameter overlay image showing concentric circle or ellipse outlines for each myelinated axon (inner ring = axon boundary, outer ring = fiber boundary). Only available in myelinated mode (not compatible with
-uor-n). The axon shape used for the overlay matches the-aargument. Output is saved with the suffix_diameter_overlay.png.
Morphometrics of a single image
Before computing the morphometrics of an image, make sure it has been segmented using AxonDeepSeg
axondeepseg_morphometrics -i test/__test_files__/__test_demo_files__/image.png -a circle -f axon_morphometrics
This generates a ‘image_axon_morphometrics.xlsx’ file in the image directory:
__test_demo_files__/
├── image.png
├── image_seg-axon.png
├── image_seg-axonmyelin.png
├── image_seg-myelin.png
├── image_axon_morphometrics.xlsx
└── pixel_size_in_micrometer.txt
Morphometrics of specific images from multiple folders
To generate morphometrics of images which are located in different folders, specify the path of the image folders using the -i argument of the CLI separated by space. For instance, to compute morphometrics of the image ‘image.png’ and ‘image_2.png’ present in the folders ‘test/__test_files__/__test_demo_files__/’ and ‘test/__test_files__/__test_segment_files__/’ respectively of the test dataset, use the following command:
axondeepseg_morphometrics -i test/__test_files__/__test_demo_files__/image.png test/__test_files__/__test_segment_files__/image_2.png
This will generate ‘image_axon_morphometrics.xlsx’ and ‘image_2_axon_morphometrics.xlsx’ files in the ‘__test_demo_files__’ and ‘__test_segment_files__’ folders:
--__test_demo_files__/
---- image.png
---- image_seg-axon.png
---- image_seg-axonmyelin.png
---- image_seg-myelin.png
---- image_axon_morphometrics.xlsx
---- pixel_size_in_micrometer.txt
...
--__test_segment_files__/
---- image_2.png
---- image_2_seg-axon.png
---- image_2_seg-axonmyelin.png
---- image_2_seg-myelin.png
---- image_2_axon_morphometrics.xlsx
---- pixel_size_in_micrometer.txt
Morphometrics of all the images present in folder(s)
To compute the morphometrics of batches of images present in folder(s), input the path of the directories using the -i argument separated by space. For example, the morphometrics files of the images present in the directory test/__test_files__/__test_segment_files__/ is computed using the following CLI command:
axondeepseg_morphometrics -i test/__test_files__/__test_demo_files__
This will generate ‘image_axon_morphometrics.xlsx’ and ‘image_2_axon_morphometrics.xlsx’ morphometrics files in the ‘test/__test_files__/__test_demo_files__’ directory:
--__test_demo_files__/
---- image.png
---- image_seg-axon.png
---- image_seg-axonmyelin.png
---- image_seg-myelin.png
---- image_axon_morphometrics.xlsx
---- image_2.png
---- image_2_seg-axon.png
---- image_2_seg-axonmyelin.png
---- image_2_seg-myelin.png
---- image_2_axon_morphometrics.xlsx
Please note that when using the axondeepseg_morphometrics command, the console output will be logged in a file called axondeepseg.log in the current working directory.
Morphometrics for Nerve Sections
You can compute morphometrics specific to nerve sections using the -n option
in the axondeepseg_morphometrics command-line interface. This enables analysis of axons
within nerve fascicle boundaries, based on a segmentation mask with the suffix _seg-nerve.png.
Currently, axondeepseg does not produce this mask, so you will need to supply it manually.
This is useful because the total nerve area allows for the calculation of axon density.
When used, the -n option performs:
Morphometric extraction of axons and myelin within the nerve boundary.
Density estimation of axons inside the fascicle.
Removal of axons located outside of the nerve mask before final metrics are saved.
axondeepseg_morphometrics -i <IMAGE_PATH> -n
The image folder must contain:
Axon mask:
*_seg-axon.pngMyelin mask:
*_seg-myelin.pngNerve segmentation mask:
*_seg-nerve.png
One output file will be generated:
<filename>_nerve_morphometrics.json: Morphometrics including axon count and density inside the nerve region.
Below is an example of the JSON file generated when using the -n option on an image with two nerve fascicles
(in other words, two disjoint regions in the _seg-nerve.png mask). This file reports the nerve fascicle areas
and their respective axon densities, as well as global area and total axon density.
{
"fascicle_areas": {
"0": {
"value": 103021.45,
"unit": "um^2",
"axon_density": {
"value": 0.00672,
"unit": "axon/um^2"
}
},
"1": {
"value": 85792.12,
"unit": "um^2",
"axon_density": {
"value": 0.00815,
"unit": "axon/um^2"
}
}
},
"total_area": {
"value": 188813.57,
"unit": "um^2"
},
"total_axon_density": {
"value": 0.00741,
"unit": "axon/um^2"
}
}
Diameter Overlay
Using the -d flag generates a grayscale PNG image (_diameter_overlay.png) showing two concentric outlines per axon: an inner ring at the axon boundary and an outer ring at the fiber (axon+myelin) boundary. This is available for myelinated axons only and works with both -a circle and -a ellipse.
axondeepseg_morphometrics -i <IMAGE_PATH> -d
axondeepseg_morphometrics -i <IMAGE_PATH> -a ellipse -d
Implementation note: The outlines are not fits to the actual mask perimeters. They are ellipses (or circles) reconstructed from the second-moment (inertia-tensor) statistics of each region, as computed by skimage.measure.regionprops (centroid, eccentricity, orientation).
For -a circle: the inner radius is axon_diam / 2 and the outer radius is axon_diam / 2 + myelin_thickness. myelin_thickness is the true uniform ring thickness (difference in equivalent radii).
For -a ellipse: axon_diam is minor_axis_length from regionprops. The inner ellipse semi-minor axis is b = axon_diam / 2 and the semi-major axis is derived as a = b / sqrt(1 - e²) from the axon eccentricity. The outer ellipse is derived independently from the axonmyelin regionprops (fiber_eccentricity, fiber_orientation).
Warning
In ellipse mode, myelin_thickness is defined as the difference in semi-minor axes between the axonmyelin and axon regions: (axonmyelin.minor_axis_length - axon.minor_axis_length) / 2. This is the myelin thickness along the minor axis only — it is not a mean or uniform perimeter thickness. A true uniform sheath on an elliptical axon does not produce an elliptical outer boundary, so the inner and outer ellipses are modeled independently from their respective regionprops statistics. For circles (-a circle), the ring is uniform and myelin_thickness has its conventional meaning.
Axon Shape: Circle vs Ellipse
Circle
Usage
axondeepseg_morphometrics -i test/__test_files__/__test_demo_files__/image.png -a circle
Studies using Circle as axon shape:
Duval et al: https://pubmed.ncbi.nlm.nih.gov/30326296/
Salini et al: https://www.frontiersin.org/articles/10.3389/fnana.2017.00129/full
Ellipse
Usage
axondeepseg_morphometrics -i test/__test_files__/__test_demo_files__/image.png -a ellipse
Studies using Ellipse as axon shape:
Bartmeyer et al: https://pubmed.ncbi.nlm.nih.gov/33542368/
Ruthig et al: https://pubmed.ncbi.nlm.nih.gov/40834043/
Note
In the literature, both equivalent diameter and ellipse minor axis are used to compute the morphometrics. Thus, depending on the usecase, the user is advised to choose axon shape accordingly.
Morphometrics file
The resulting ‘axon_morphometrics.csv/xlsx’ file will contain the following columns headings. Most of the metrics are computed using skimage.measure.regionprops.
By default for axon shape, that is, circle, the equivalent diameter is used. For ellipse axon shape, minor axis is used as the diameter. The equivalent diameter is defined as the diameter of a circle with the same area as the region.
Field |
Description |
|---|---|
x0 |
Axon X centroid position in pixels. |
y0 |
Axon Y centroid position in pixels. |
gratio |
Ratio between the axon diameter and the axon+myelin (fiber) diameter (gratio = axon_diameter / axonmyelin_diameter). |
axon_area |
Area of the axon region in \({\mu}\)m2. |
axon_perimeter |
Perimeter of the axon object in \({\mu}\)m. |
myelin_area |
Difference between axon+myelin (fiber) area and axon area in \({\mu}\)m2. |
axon_diameter |
Diameter of the axon in \({\mu}\)m. |
myelin_thickness |
Half of the difference between the axon+myelin (fiber) diameter and the axon diameter in \({\mu}\)m. |
axonmyelin_area |
Area of the axon+myelin (fiber) region in \({\mu}\)m2. |
axonmyelin_perimeter |
Perimeter of the axon+myelin (fiber) object in \({\mu}\)m. |
solidity |
Ratio of pixels in the axon region to pixels of the convex hull image. |
eccentricity |
Eccentricity of the ellipse that has the same second-moments as the axon region. |
orientation |
Angle between the 0th axis (rows) and the major axis of the ellipse that has the same second moments as the axon region. |
fiber_eccentricity |
Eccentricity of the ellipse that has the same second-moments as the axon+myelin (fiber) region. |
fiber_orientation |
Angle between the 0th axis (rows) and the major axis of the ellipse that has the same second moments as the axon+myelin (fiber) region. |
image_border_touching |
Flag indicating if the axonmyelin objects touches the image border |
bbox_min_y |
Minimum y value of the bounding box (in pixels). This bound is inclusive. |
bbox_min_x |
Minimum x value of the bounding box (in pixels). This bound is inclusive. |
bbox_max_y |
Maximum y value of the bounding box (in pixels). This bound is exclusive. |
bbox_max_x |
Maximum x value of the bounding box (in pixels). This bound is exclusive. |
Colorization & Instance Segmentation
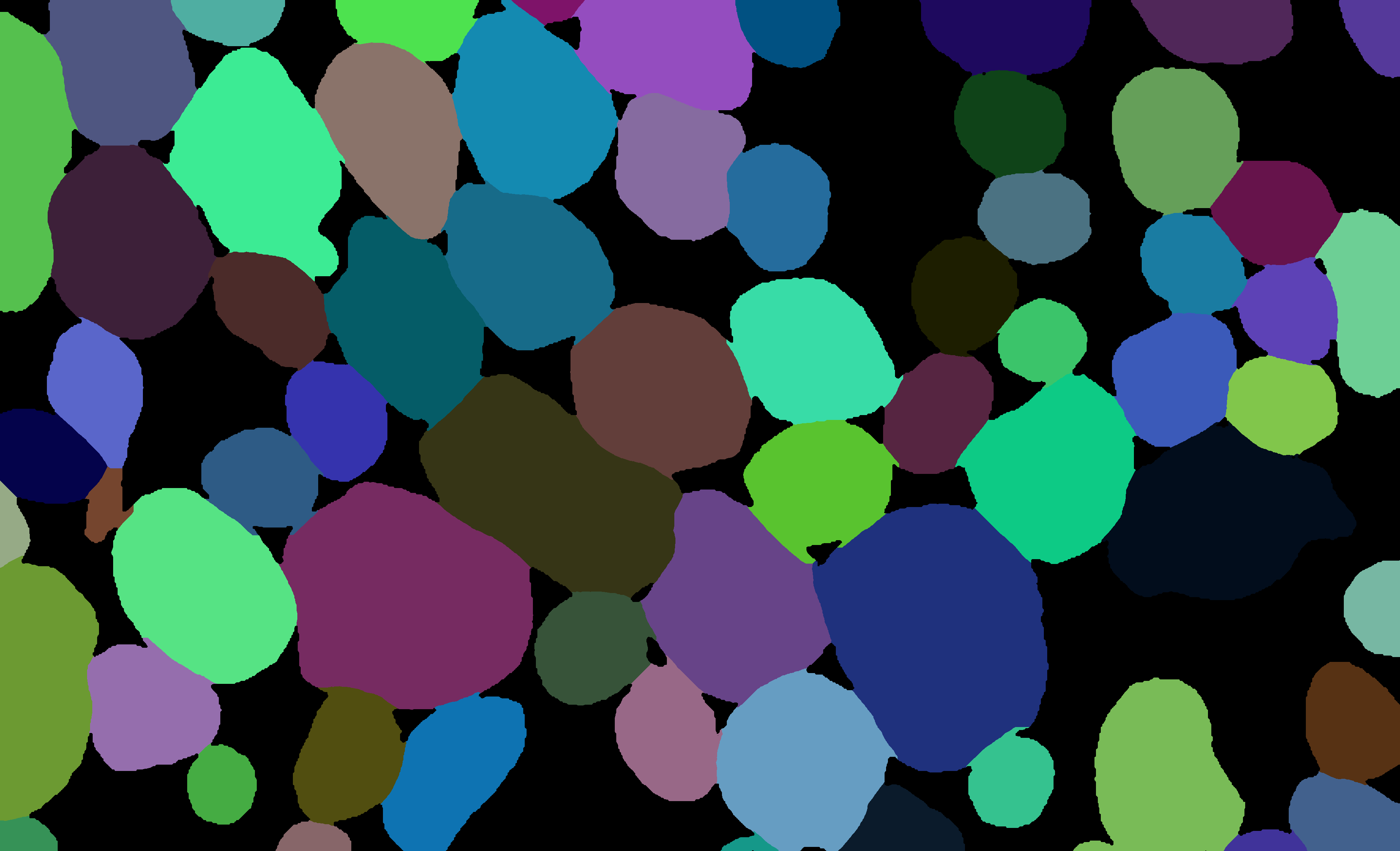
During the morphometrics computation, axondeepseg internally converts the semantic segmentation (output of the deep learning model) into an instance segmentation. This step is essential to take measurements on individual axons when the axon density is high, because if two or more elements have their myelin touching, the software needs to know which axon it is attached to. Using the -c flag, you can obtain the colorized instance segmentation to take a look at this internal representation. The image below illustrates what a typical instance segmentation looks like. This option will also save the raw instance segmentation in 16-bit format (with value 0 for background, 1 for the first axon, 2 for the second axon, etc.) in the same folder as the input image, with the suffix _seg-instance-map.png.

Implementation details
The following sections provide more details about the implementation of the algorithms behind the morphometrics computation.
Diameter estimation
The diameter \(D\) is computed differently based on the chosen axon shape:
For the circle axon shape, the diameter is simply the equivalent diameter of the axon region, which is the diameter of a circle with the same area as the axon region.
For the ellipse axon shape, the computation is entirely different. We do not actually need to fit an ellipse to get the minor axis length. Instead,
sklearncomputes this by using the second order central moments of the image region, which represents the spatial covariance matrix of the image. By computing its eigenvalues, we get the moment of inertia along the axis with the most variation and the axis with the least variation, which are respectively the major and minor axes of the ellipse. We can recover the minor axis length using the moment of inertia formula:\[I = \frac{1}{4} mr^2 \Leftrightarrow r = 2\sqrt{\frac{I}{m}}\]Assuming a uniform unit mass, we finally get \(D = 2r = 4\sqrt{I}\).
Eccentricity estimation
The eccentricity computation is based on the same principle as the diameter estimation for the ellipse axon shape. We use the eigenvalues of the second order central moment of the image, which gives us the moment of inertia along the major axis and the minor axis. The formula to compute the eccentricity of an ellipse is \(e = \sqrt{1 - \frac{b^2}{a^2}}\), where \(a\) and \(b\) respectively represent the lengths of the semi-major and semi-minor axes. Since the ratio \(\frac{a}{b}\) is equivalent to the ratio of the central moment eigenvalues, they are used instead of the actual lengths because they are easier to compute.
Postprocessing
The morphometrics computation can be followed by optional postprocessing steps. The following sections describe the postprocessing options available in AxonDeepSeg.
Morphometrics aggregation
This feature aggregates morphometrics per subject. It is useful when you have multiple images per subject and you want to compute statistics per subject.
To use this feature, you need to have done the following:
Segment all the images of interest with AxonDeepSeg using the axondeepseg command.
Compute the morphometrics of all the images using the axondeepseg_morphometrics command. Make sure that the morphometrics files are named with the same prefix as the image name (e.g. image_axon_morphometrics.xlsx for image.png).
Have the following folder structure:
folder_with_all_subjects/ ├── subject1/ │ ├── image1.png │ ├── image1_axon_morphometrics.xlsx │ ├── image2.png │ ├── image2_axon_morphometrics.xlsx │ ├── ... ├── subject2/ | ├── ... └── ...
To aggregate the morphometrics per subject, use the following command:
axondeepseg_aggregate -i folder_with_all_subjects
This will generate a folder called morphometrics_agg in the input folder, containing the aggregated morphometrics per subject. It will also contain a short summary file named statistics_per_axon_caliber.xlsx which contains basic statistics for axon diameter, myelin thickness and g-ratio. These statistics are computed per axon diameter range.
Quality Assessment (QA) Report
AxonDeepSeg now includes a comprehensive Quality Assessment (QA) feature that generates interactive HTML reports for evaluating segmentation results. This feature provides detailed visualizations and statistics to help users assess the quality of their axon and myelin segmentations.
Generating QA Reports
Through Napari Plugin
The QA report can be generated directly from the AxonDeepSeg Napari plugin:
Open Napari and the AxonDeepSeg plugin.
Load your image.
Load your mask or generate a segmentation using the plugin.
Generate a Morphometrics file using the plugin.
Click the “QA Report” button in the plugin interface
The report will automatically generate and open in your default web browser
The report is saved in a QA folder within your morphometrics directory, containing:
Interactive HTML report (
AxonDeepSeg_QA_Report.html)Individual metric histograms
Segmentation overlays
Individual axon closeup images
Report Features
The QA report includes several interactive sections:
Summary Section
Key Statistics: Mean ± standard deviation for axon diameter, myelin thickness, and g-ratio
Segmentation Visualization: Overlay of axon (blue) and myelin (red) masks on the original image
Interactive Toggle: Press and hold ‘T’ to temporarily view the original image without segmentation overlays
Histograms Section
Distribution Analysis: Interactive histogram viewer for the mophrometrics.
Available Metrics: Axon diameter, myelin thickness, g-ratio, axon area, myelin area
Statistical Summary: Each histogram displays mean, median, standard deviation, IQR, min/max values, and NaN counts
Single Axon Viewer
Individual Axon Inspection: Navigate through each axon with detailed closeup views
Dual Image Display: Toggle between original and labeled views for each axon
Metric Rankings: Percentile rankings and absolute ranks for diameter, thickness, and g-ratio
Flagging System: Manually flag axons as:
Good Segmentation (Green) - Press ‘1’
Needs Correction (Yellow) - Press ‘2’
Marked for Deletion (Red) - Press ‘3’
Clear Flag - Press ‘0’
Keyboard Shortcuts
Function |
Shortcut |
|---|---|
Previous axon |
Left Arrow (←) |
Next axon |
Right Arrow (→) |
Show original (hold) |
T key |
Good segmentation |
1 key |
Needs correction |
2 key |
Mark for deletion |
3 key |
Clear flag |
0 key |
Export Features
Export Flagged Axons button: Export a CSV file containing all flagged axons with their IDs, status, diameter, and g-ratio.
Technical Details
The QA report generation process:
Morphometrics Analysis: Uses the existing morphometrics computation pipeline
Image Processing: Generates individual axon closeups with buffer regions around each axon
Statistical Analysis: Computes comprehensive statistics for all morphometric parameters
Interactive Visualization: Creates an HTML-based report with JavaScript-powered interactivity
The report includes:
Sample identification and generation timestamp
Software version information
Comprehensive metric distributions
Individual axon inspection capabilities
Professional styling and responsive design
The QA feature is particularly useful for:
Quality Control: Quickly identifying segmentation issues
Research Validation: Statistical validation of segmentation results
Manual Correction: Targeted identification of axons needing manual intervention
Result Documentation: Professional reporting for publications and presentations
Manual correction of segmentation masks
If the segmentation with AxonDeepSeg does not give optimal results, you can try one of the following options:
Napari plugin
Open image and mask
Open Napari by entering ads_napari in the terminal.
Load the AxonDeepSeg plugin using the Napari toolbar: Plugins -> ADS (AxonDeepSeg)
Load the microscopy image using the Napari toolbar: File -> Open file(s)
- If no segmentation masks already exists:
Choose one of AxonDeepSeg’s default models in the dropdown menu “Select the model”
Then click on the “Segment image” button
- If a segmentation masks already exists:
Click on the “Load mask” button and select the image with the suffix “_seg-axonmyelin”
After a mask is loaded or generated, the axon (blue) and myelin (red) layers will be overlayed on top of the histology image.
To pan on the image, click and drag your trackpad or mouse.
Modify the mask
Click the mask (myelin or axon) that you want to modify in the “layer list” panel.
To edit the mask you chose, select one of the three editing modes in the “layer control” panel on the left.
Note
Zooming and panning are disabled while editing the mask. To regain these functionalities, click on the magnifying icon  to re-activate it.
to re-activate it.
The “Fill axons” button in the AxonDeepSeg plugin (right panel) can also be used to edit the masks, and overall can speed up your workflow.
Note
The “Fill axon” button will fill closed myelin mask areas by painting in the axon mask. A good workflow if starting from scratch would be to manually segment all the myelin in the image and then click the “Fill axons” button to fill in the axon areas.
Warning
The “Fill axons” functionality will not behave properly if there are myelin objects not closed, or if multiple myelin objects touch each other to form a big closed cluster.
Axon removal toggle
Note
To use this feature, the image must have been loaded after the plugin, and the mask must have been loaded or generated. Lastly, the image layer must be selected.
The “Axon removal toggle” button in the AxonDeepSeg plugin (right panel) can be used to remove axons from the mask.
After activated, CTRL-click on the axon you want to remove. The axon will be removed from the mask.
Note
To undo, select the axon layer and press CTRL+Z, then repeat with the myelin mask.
To deactivate the “Axon removal toggle” mode, click on the button again.
Show axon metrics
Note
To use this feature, the image must have been loaded after the plugin, and the mask must have been loaded or generated. Lastly, the image layer must be selected.
The “Show axon metrics” button in the AxonDeepSeg plugin (right panel) can be used to view some key metrics of a clicked axon.
After activated, ALT/OPTION-click on the axon you want to view the metrics of. A pop-up window will show the metrics of the clicked axon at the bottom right of the image viewer.
To deactivate the “Toggle axon removal” mode, click on the button again.
Save the mask
Click the “Save segmentation” button in the AxonDeepSeg plugin (right panel).
Note: In case of an overlap between the axons mask and the myelin mask, the myelin will have priority when saving the new segmentation.
The “_seg-axon.png” and “_seg-myelin.png” are the axons-only and myelin-only binary masks.
- The “_seg-axonmyelin.png” file is the axon+myelin mask.
Note that this mask is a PNG 8-bit file with 1 channel (256 grayscale), with color values of 0 for background, 127 for myelin and 255 for axons.
GIMP software
To create a new axon+myelin manual mask or to make manual correction on an existing segmentation mask, you can use the GIMP software (Link for download).
If you are making correction on an existing segmentation mask, note that when you launch a segmentation, in the folder output, you will also find the axon and myelin masks (with the suffixes ‘_seg-axon.png’ and ‘_seg-myelin.png’). You can then manually correct the myelin mask and create a corrected axon+myelin mask.
For a detailed procedure, please consult the following link: Manual labelling with GIMP.
Running the testing suite
Developers are encouraged to run the testing suite in the terminal, and develop relevant tests as needed.
To run the testing suite, run the following command within the AxonDeepSeg directory:
axondeepseg_test --full
Alternatively, pytest commands can be run directly from the terminal for more control over the testing suite. For example, to run all tests, use:
pytest test/
and to run a specific test file, use:
pytest test/test_segment.py
Citation
If you use this work in your research, please cite:
Zaimi, A., Wabartha, M., Herman, V., Antonsanti, P.-L., Perone, C. S., & Cohen-Adad, J. (2018). AxonDeepSeg: automatic axon and myelin segmentation from microscopy data using convolutional neural networks. Scientific Reports, 8(1), 3816. Link to the paper.